
Kertesz. 'Israel has an abundance of talent and motivation, but not of biotech investments. [Biotech] requires tens of millions of dollars and years of development, which aren’t always fruitful.' Photo by Eyal Toueg
A small American start-up managed to streamline complex DNA-sequencing techniques. Its co-founder, Mickey Kertesz, thinks the greatest breakthrough may come in the field of cancer treatment.
Early last month, several dozen scientists were invited to attend a conference in San Diego, California, on the genomic revolution. Instead of receiving a wireless mouse or a T-shirt with a witty slogan, each attendee was awarded his own personal genome sequence: a breakdown of all the genetic material he carries in his body. The term “awarded” is a bit misleading, since each person paid $5,000 for this information (which also included the tablet computer on which the genome breakdown was downloaded). But compared to the $2.7 billion that was invested in mapping the first human genome 10 years ago, and considering that only a few thousand people in the entire world know their genome sequence − you could certainly say that was a good deal.
That was largely the message the conference, entitled “Understand Your Genome," tried to get across: What was referred to as the Human Genome Project has in effect lost its capital letters, and very soon will become an everyday product. Hereafter it will be translated into the individual genomes of the proverbial Riki Cohen of Hadera or John Smith of Nebraska. Or as a professor from the Stanford University School of Medicine who attended the recent California gathering put it: “Every day, I drive past this Ferrari dealership in Palo Alto and I see the 458 Ferrari Spider, which retails at $398,000. I’ve worked out that if that had been the cost of sequencing at the time of the Human Genome Project and the price had dropped at the same rate, the car would now cost 40 cents.”
The organizer of the conference, the American company Illumina, has good reason to celebrate the genomic revolution. Illumina already has a monopoly on genetic sequencing: Ninety percent of the genetic data obtained in the world today is generated by means of its technologies. Furthermore, late last year it acquired a small and unusual startup called Moleculo, and thereby clinched one of the more interesting deals in the global biotech industry: After less than a year’s work, at an investment of less than $1 million (and with a name that was borrowed from a Conan O’Brien parody of Superman on “Saturday Night Live”), Moleculo propelled DNA sequencing to new heights of efficiency.
One of the startup’s two co-founders is an Israeli, Dr. Michael (“Mickey”) Kertesz, 42, who was in the midst of research at Stanford when he found himself in the eye of a commercial storm. He speaks about Moleculo’s achievement modestly, but confidently, as something that will change the battlefield on which we combat diseases such as cancer, AIDS and even depression.
“Instead of shooting in the dark, as is the case today,” he said in a telephone interview, “we will be able to rely on precise data that will enable us to attack targets in a focused manner, and therefore effectively.”
It is perhaps not surprising that Kertesz uses such imagery, considering that he served as a pilot in the Israel Air Force. After his discharge, he went to study computer science, delighted in the start-up years of the late 1990s, and charted a long and stable relationship with the high-tech industry, until he encountered the field of biology.
“I even remember the moment I fell in love with life sciences,” he says. “It happened on a paragliding trip in Australia. I was sitting in the middle of nature and reading Darwin’s ‘The Origin of Species,’ and every once in a while I found myself lifting my head, looking around me and beginning to understand the world differently. It was a very powerful sensation, which recurred later, when I started reading biology texts and understanding the molecular mechanisms that are at the basis of evolution. I remember the amazement. It astounded me to think how at any given moment, complex processes are taking place in our bodies at an insane pace and with fantastic precision. Thousands of little machines, operating at an almost inconceivable degree of sophistication and accuracy.

Moleculo laboratory staff. Photo by Eyal Toueg
“I was working in high-tech at the time, and biology was an exciting hobby, but at a certain stage, it was no longer enough for me. I decided to turn the hobby into a career, and I completed a doctorate in computational biology at the Weizmann Institute [of Science]. The obvious next step was to do a post-doctorate, and that is how I came to Stanford, to a lab that studies viruses.
“In the course of the research, I tried to monitor the rapid change of viruses and to discern the tiny differences in their genome, but existing DNA sequencing methods were not accurate enough for my research. A Russian doctoral student named Dmitry Pushkarev, who worked in the lab at that time, also encountered this problem from a completely different direction in his research. Together we tried to develop ways of making the genome sequencing process more precise and effective. At first it was in order to solve our personal [research] issues, but we quickly realized that the solution could have many other applications. In late 2011, we founded the company, Moleculo, to commercialize the method and provide rapid access to our technology.”
What is your innovation?
“When molecular biology was getting started, determining a DNA sequence was a heroic act. It was a lengthy, manual process which could be applied only to a limited number of DNA segments. The Human Genome Project, which purported to sequence the entire genome, gave the technology a tremendous push and made it nearly automatic. In the past five years, companies such as Illumina have developed machines that we can describe as receiving a DNA sample at one end and spitting out a file containing its sequence at the other end, at a speed that’s thousands of times faster than ever before.
“Nevertheless, despite the sophisticated process, one obstacle remained. If we use the familiar metaphor by which the genome is described as a book and the bases of DNA are the letters that make up the text − we can say that the machines cannot read all of the text at once but rather only in sentences of 100 letters. That is problematic when it comes to our genome, for example, which has 3 billion letters, but also in cases where less lengthy genomes are concerned.
“To overcome this obstacle, you do the following: Shred the book into little pieces, each containing a sentence of 100 letters, and give them to the machine to read. You then connect the short sentences to each other, thus reconstructing the original sequence. In principle, the sentence order can be determined because of the fact that the end of one sentence appears at the beginning of the sentence next to it. But unfortunately, the genomic text contains a great many segments that repeat themselves in various places. For this reason, sentences of 100 letters are not unique enough to determine with certainty where they should correctly be placed within this enormous text. This recurrence introduces errors into the reconstruction process, and we are after accuracy, after all: In order to understand the meaning of a text, and to discover when and where a particular sentence in it became corrupted, the reading must be precise.
“And this is where our development comes in. To carry on with the book metaphor, instead of shredding the book immediately, we first dye its pages, each page with a different color. Then we shred the book and give it to the machine to read. The machine still knows how to read only 100-letter sentences, but this time they are dyed the color of the page from which they were taken. In this way, our system can deal with each page separately − red sentences apart, blue apart, and so on. This makes the task of arranging the sentences into a sequence much simpler. Now it is clear that all of the sentences dyed the same color came from the same page. In other words, the process can be divided into two stages: First, reconstructing each page separately with the help of the colored sentences, and then finding the order within the pages. It makes the process a lot more efficient and less prone to mistakes.”
Fewer puzzle pieces
How is genomic text dyed?
“Through molecular marking. We attach a different molecular label to each DNA segment that is 10,000 bases long. To use a different metaphor, that of a puzzle, we can say that instead of putting together a puzzle of 100,000 little pieces, now we have to put together 10 big pieces. That is much easier, of course.”
In other words, your technology is based on a trick from the field of molecular biology?
“The technology we developed includes a process that is based on molecular biology − of dyeing − and also a computational component from the field of bioinformatics, which makes it possible to handle the new type of data you get. Each of the stages is based on existing methods, but the fact that they are combined, along with the reciprocal relations between them has brought about significant results.”
What breakthroughs might occur thanks to your development?
“In addition to the possibility of handling very complicated genomes, such as those of plants, which are typically far more complex than those of animals, the availability of hereditary material that can be read contains promise for everything related to personalized medicine. It accelerates the possibility of treating a patient in a manner that takes into account the genetics unique to him, the particular stage of his disease, and his sensitivity to medication and other treatments. In my opinion, the greatest breakthrough will be in the field of cancer treatment.”
As in the case of Angelina Jolie?
“Jolie’s story made headlines touting our ability to determine statistically what a woman’s odds are of developing breast cancer by determining segments in her genome. But we are talking about much more than this. Today every single patient with a particular cancer, for example lung cancer, is given similar treatment. That is a very rough approach, because every tumor is different. Remember that a tumor is made of cells whose genome underwent a change, which turned them cancerous. Every cancerous tumor has a characteristic and unique genome: This genome is different from that of the rest of the patient’s cells, it is different from one patient to another, and is not even similar to what it was itself a few months ago. This is because cancer is a disease that develops over time, in parallel to changes in the tumor genome.
“Returning to the book metaphor: In the text of a certain cancerous tumor, there can be 50 pages that appear twice, 50 other pages can be missing, and over time additional changes can occur. It is a very disorderly genome. Any change in the genome causes the disease to be a little different and to respond to treatments differently. The possibility of establishing the cancerous genome sequence easily and precisely will allow us to identify all of these changes, monitor them throughout the disease, and adjust the right medication for them or any other treatment. It could be a genuine revolution.”
What about other fields?
“At the Personalized Medicine World Conference held recently in California, I heard a lecture by a colleague at a firm that focuses on mental illnesses such as depression. I was amazed. Today, a person who suffers from depression receives a particular medication without any knowledge of the extent of its compatibility for him. It takes weeks to begin taking effect, and if it turns out to be ineffective, he tries another random medication, and once again endures weeks of suffering.
“To prevent this vague process of trial and error, the company tested thousands of people, who responded in various ways to various medications, and found a correlation between unique segments in the patients’ genome and the success of certain treatments for depression. This way, when a person with depression comes to the company, it sequences parts of his genome and can predict the probability that a particular treatment will work. In other words, it is possible to know ahead of time which medication might help him, if at all.”
Early exit
Your company was acquired within a remarkably short time and after an incredibly small investment. How did that happen?
“First of all, it is important to understand that we did not develop a new machine for establishing the genome sequence. Such a task takes hundreds of millions of dollars and many years of development. Our development takes advantage of existing sequencing technologies, which we enhanced by altering the DNA before it enters the machine and by developing a computational stage that analyzes the result by a unique method.
“Second, we were careful not to blow the company up beyond what was necessary for promoting the technology, and we managed to create an income source at a very early stage. Our technology was operational after just four months, and what we did subsequently was to turn it into a commercial product. Here we were in for a surprise. While we were dealing with development, word about us got around, and clients began approaching us. The market was thirsting after a technology like this, and more and more companies and institutions approached us with requests to use the new technology. They were prepared to pay, and even though we did not have a finished product yet, we were able to offer them a service for sequencing genomes at a higher quality than ever before.
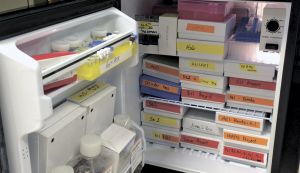
Refrigerator in which the DNA samples are carefully preserved. Photo by Eyal Toueg
“That is how we got to work with clients like the U.S. Department of Energy and Department of Agriculture, which are interested in playing with the genome of plants to turn them into biofuel, and with research labs at Stanford, Harvard, Berkeley and more. They sent us their DNA samples, and we used our methods to provide them with long and accurate sequences. In addition to the money these clients brought in, they provided us with important feedback that helped us improve the product in real-time.
“We got advise from excellent outside consultants, who helped us a lot with marketing and business development, and of course we also had a lot of luck. But beyond that, it seems to me the thing is that we simply offered a solution to a real problem. Both Dmitry and I identified the need for long reads from our own research questions, and other people identified this need to consider other questions. Ultimately, it seems we possessed a unique technology that met a genuine need.”
What about you? Are you staying in biotech or going back to academia?
“For now I and the rest of the team are working at Illumina on the development and assimilation of the technology in preparation for launching it. But my passion is being at the place where new things are happening. I always prefer the revolutionary stage to the establishment stage that follows, so I presume that at a certain point I will seek out new challenges. I would like to continue my virus research, which was suspended because of setting up the company. Our children and grandchildren won’t believe it when we tell them that once upon a time, when a virus attacked our body, we would be sent to bed with a cup of tea. It is intolerable in my opinion that millions of people die every year from AIDS, hepatitis and influenza viruses, and I believe that is about to change.
“The problem with diseases like AIDS and influenza is that the genome of the viruses that generates them undergoes mutations at a tremendous pace. Therefore, a patient carries within him not a single strain, but rather an entire population of viruses. The ability to monitor its change with maximum speed and precision might cause a paradigm shift in our approach to viruses. The possibility of mapping the genomes of viruses will move the war against them from the cellular battlefield − where for example you look at the virus with a microscope, examine what it’s attached to and study its proteins − to a digital-genomic battlefield, which is based on a huge amount of genomic data. This way you can assess which genomic domain the virus prefers, in which domain it cannot live, and which medications can affect all of these.”
Basically, you are saying that biology is going to become a more exact science?
“Precisely. That is the paradigm shift I’m talking about. The possibility of exercising a mathematical-computational approach on biology will transform virus research and medical treatment of viruses into much more quantitative and exact. Personally, I would like to be a part of that.”
In Israel?
“Certainly. I still don’t know when, but that is obvious. I think the message is clear: The Jewish mind was always good at original ideas, and Israel has an abundance of talent and motivation, but not as much biotech investments. The thing is that biotech has a reputation today of being a demanding industry that requires tens of millions of dollars and years of development, which are not always fruitful.
“I would be delighted if the story of Moleculo and other companies like ours can change something in the attitude toward biotech, and inspire both investors and entrepreneurs. It seems to me that this kind of entrepreneurship, based on developing technology that is essentially an innovative combination of existing components, may be suitable for the Israeli ecosystem.”
The sale price for Illumina was never published. I assume you earned more than gratification from it.
“The sale happened pretty fast. We were approached by quite a few companies that had heard about us from our clients and wanted to collaborate with us or to gain access to the product. Negotiations with Illumina moved pretty fast. At the start of the talks with them we focused on access to their marketing and production setup, but after a few weeks of working together, they showed up with an offer to buy the company. We felt it could be an excellent home for the technology, and after brief negotiations we reached a deal that was fair to all the parties. That was an insane time, because the final signing of the acquisition deal took place two days before we were supposed to close a venture capital financing round. As in other cases, the acquiring company decided not to publicize the financial details of the deal, and we respect that.”
What does the family say?
“A process of scientific research is always accompanied by ups and downs: One day the experiment works, and the next day it suddenly looks like everything is falling apart. Add to that the ups and downs related to the business side of managing a company and you will understand what a wild period that was. My wife Tali supported me throughout and there is no way I could have survived without her support.
“My daughters, aged 5 and 7, are very interested in this subject and love to visit my lab. The stories they like best to hear before bed are about DNA. I sit by their bedsides and tell them, for example, about the struggle between viruses and the immune system. Nature provides me with countless fascinating stories about a spaceship-like virus that attaches to the cell and injects it with a DNA chain, or about cells of the immune system that swallow other cells. They are sure it’s a fantasy, a children’s story, but it is all the absolute truth. I go on and on until they fall asleep. And sometimes, after an exhausting day’s work, I fall asleep along with them.”
